Handling mothur data
Calculate relative abundances
You can read a shared file and calculate relative abundances with
calc_relabun():
shared_dat <- read_tsv(system.file("extdata", "test.shared", package = "schtools"))
relabun_dat <- shared_dat %>%
calc_relabun()
head(relabun_dat)
#> # A tibble: 6 × 3
#> sample otu rel_abun
#> <chr> <chr> <dbl>
#> 1 p1 Otu0001 0
#> 2 p1 Otu0003 0
#> 3 p1 Otu0004 0
#> 4 p1 Otu00008 0
#> 5 p1 Otu0044 0.25
#> 6 p1 Otu0056 0.25calc_relabun() returns the data frame in long format.
You can use tidyr::pivot_wider() to convert it to wide
format:
wide_dat <- relabun_dat %>%
pivot_wider(names_from = "otu", values_from = "rel_abun")
head(wide_dat)
#> # A tibble: 6 × 13
#> sample Otu0001 Otu0003 Otu0004 Otu00008 Otu0044 Otu0056 Otu0057 Otu0058
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 p1 0 0 0 0 0.25 0.25 0 0.25
#> 2 p2 0.167 0 0.167 0 0.167 0 0 0.167
#> 3 p3 0 0.2 0 0 0.2 0 0 0
#> 4 p4 0.25 0.25 0 0.25 0 0 0 0
#> 5 p5 0.25 0 0.25 0 0.25 0 0 0
#> 6 p6 0.167 0 0 0.167 0 0.167 0.167 0.167
#> # ℹ 4 more variables: Otu0159 <dbl>, Otu0208 <dbl>, Otu0328 <dbl>,
#> # Otu0329 <dbl>You can see that the relative abundances for each sample sum to 1:
wide_dat %>%
select(starts_with("Otu")) %>%
rowSums()
#> [1] 1 1 1 1 1 1 1 1 1 1Taxonomy files
mothur formats taxonomy files as tab-separated values (tsv). You can
use read_tax() to parse the taxonomy data and create
separate columns for each taxonomic level.
tax_dat <- read_tax(system.file("extdata", "test.taxonomy", package = "schtools"))
head(tax_dat)
#> # A tibble: 6 × 10
#> otu otu_label tax_otu_label label_html kingdom phylum class order family
#> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
#> 1 Otu0001 OTU 1 Bacteroides (… <i>Bacter… Bacter… Bacte… Bact… Bact… Bacte…
#> 2 Otu0003 OTU 3 Porphyromonad… <i>Porphy… Bacter… Bacte… Bact… Bact… Porph…
#> 3 Otu0004 OTU 4 Porphyromonad… <i>Porphy… Bacter… Bacte… Bact… Bact… Porph…
#> 4 Otu00008 OTU 8 Enterobacteri… <i>Entero… Bacter… Prote… Gamm… Ente… Enter…
#> 5 Otu0044 OTU 44 Bacteria (OTU… <i>Bacter… Bacter… Bacte… Bact… Bact… Bacte…
#> 6 Otu0056 OTU 56 Bacteria (OTU… <i>Bacter… Bacter… Bacte… Bact… Bact… Bacte…
#> # ℹ 1 more variable: genus <chr>The column label_html provides html that correctly
italicizes the genus name without italicizing the OTU label. This can be
used with ggtext::element_markdown() to make nice
plots:
library(ggtext)
set.seed(20220427)
relabun_dat %>%
mutate(sample_num = stringr::str_remove(sample, "p") %>%
as.integer(), treatment = case_when(sample_num%%2 == 1 ~ "A", TRUE ~ "B")) %>%
inner_join(tax_dat, by = "otu") %>%
ggplot(aes(x = rel_abun, y = label_html, color = treatment)) + geom_jitter(alpha = 0.7,
height = 0.2) + labs(x = "Relative abundance", y = "") + theme_minimal() + theme(axis.text.y = element_markdown())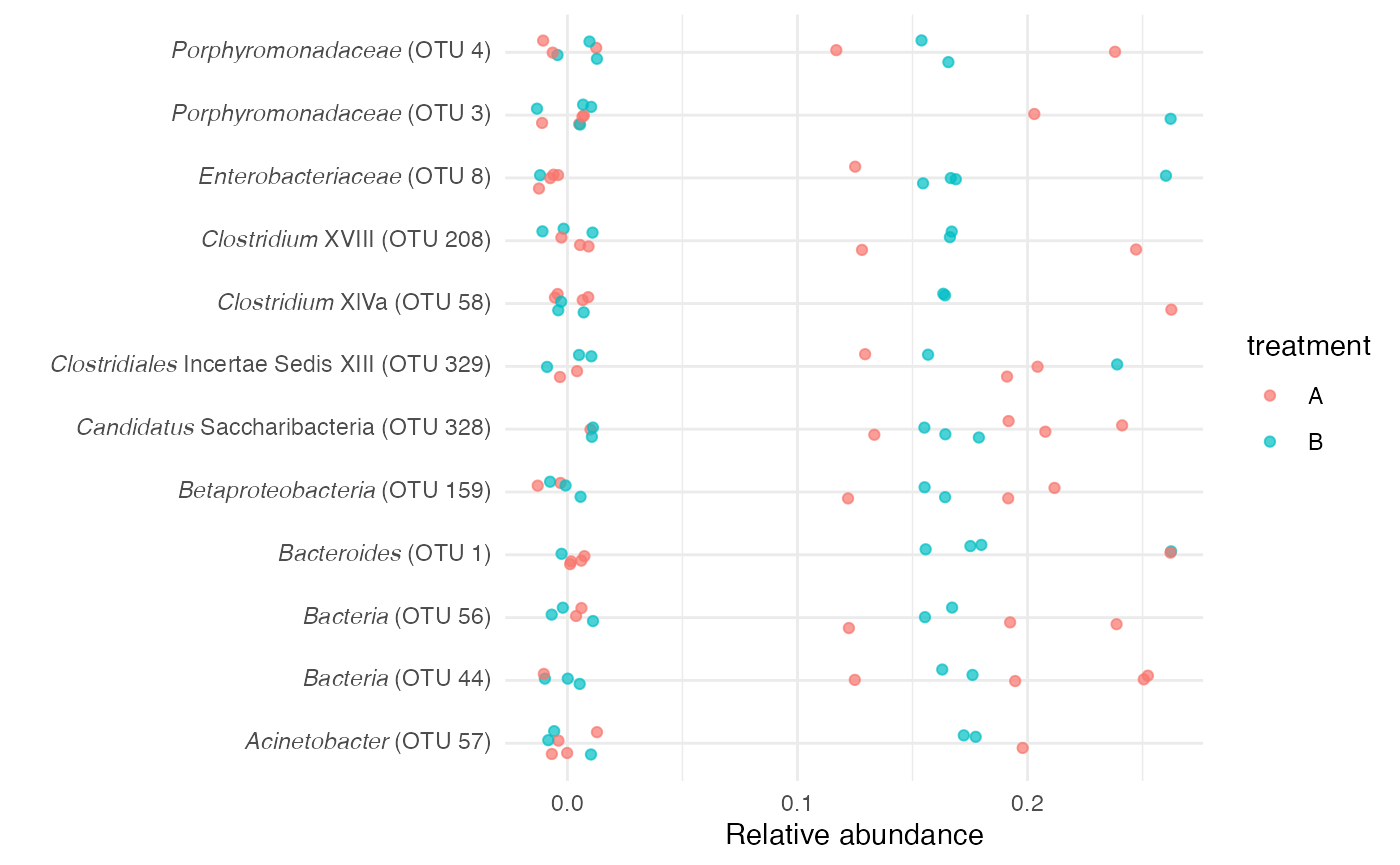
Pooling OTU counts at different taxonomic levels
A common task is to repeat OTU-level analyses at different taxonomic levels to determine which resolution is optimal for answering your questions. You’ll need a shared file, generated from clustering sequences into OTUs with mothur, and a corresponding taxonomy file. Take a look at the mothur documentation for info on generating these files and performing microbiome analyses.
In this example, pool_taxon_counts() pools the OTU
counts in the shared file at the genus level and returns new shared and
taxonomy data frames.
tax_dat <- read_tax(system.file("extdata", "test.taxonomy", package = "schtools"))
shared_dat <- readr::read_tsv(system.file("extdata", "test.shared", package = "schtools"))
pool_taxon_counts(shared_dat, tax_dat, "genus")
#> $shared
#> # A tibble: 10 × 13
#> label Group numOtus Otu01 Otu02 Otu03 Otu04 Otu05 Otu06 Otu07 Otu08 Otu09
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 genus p1 10 0 0 0 2 0 1 0 1 0
#> 2 genus p10 10 1 0 1 0 1 0 1 1 1
#> 3 genus p2 10 1 1 0 1 0 1 0 0 1
#> 4 genus p3 10 0 1 0 1 0 0 1 0 1
#> 5 genus p4 10 1 1 1 0 0 0 0 0 0
#> 6 genus p5 10 1 1 0 1 0 0 0 0 1
#> 7 genus p6 10 1 0 1 1 1 1 0 0 1
#> 8 genus p7 10 0 0 0 1 1 0 1 0 1
#> 9 genus p8 10 0 1 1 2 0 0 1 1 0
#> 10 genus p9 10 0 1 1 2 0 0 1 1 1
#> # ℹ 1 more variable: Otu10 <dbl>
#>
#> $tax
#> # A tibble: 10 × 3
#> otu size genus
#> <chr> <dbl> <chr>
#> 1 Otu01 5 Bacteroides
#> 2 Otu02 6 Porphyromonadaceae unclassified
#> 3 Otu03 5 Enterobacteriaceae unclassified
#> 4 Otu04 11 Bacteria unclassified
#> 5 Otu05 3 Acinetobacter
#> 6 Otu06 3 Clostridium XlVa
#> 7 Otu07 5 Betaproteobacteria unclassified
#> 8 Otu08 4 Clostridium XVIII
#> 9 Otu09 7 Candidatus Saccharibacteria unclassified
#> 10 Otu10 5 Clostridiales Incertae Sedis XIII unclassifiedYou can do this for any taxonomic level in your taxonomy data frame.
pool_taxon_counts(shared_dat, tax_dat, "phylum")
#> $shared
#> # A tibble: 10 × 8
#> label Group numOtus Otu1 Otu2 Otu3 Otu4 Otu5
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 phylum p1 5 0 0 2 2 0
#> 2 phylum p10 5 1 3 0 1 1
#> 3 phylum p2 5 2 0 1 2 1
#> 4 phylum p3 5 1 1 1 1 1
#> 5 phylum p4 5 2 1 0 1 0
#> 6 phylum p5 5 2 0 1 0 1
#> 7 phylum p6 5 1 2 1 1 1
#> 8 phylum p7 5 0 2 1 1 1
#> 9 phylum p8 5 1 2 2 1 0
#> 10 phylum p9 5 1 2 2 2 1
#>
#> $tax
#> # A tibble: 5 × 3
#> otu size phylum
#> <chr> <dbl> <chr>
#> 1 Otu1 11 Bacteroidetes
#> 2 Otu2 13 Proteobacteria
#> 3 Otu3 11 Bacteria unclassified
#> 4 Otu4 12 Firmicutes
#> 5 Otu5 7 Candidatus SaccharibacteriaDistance files
If you have a distance file saved as a phylip-formatted lower
triangle matrix from mothur’s dist.seqs
command, you can read it into R with read_dist():
dist_filepath <- system.file("extdata", "sample.final.thetayc.0.03.lt.ave.dist",
package = "schtools")
dist_tbl <- read_dist(dist_filepath)
head(dist_tbl)
#> # A tibble: 6 × 3
#> rows columns distances
#> <chr> <chr> <dbl>
#> 1 104_1_D1 104_1_D0 0.893
#> 2 104_1_D10 104_1_D0 0.254
#> 3 104_1_D10 104_1_D1 0.922
#> 4 104_1_D2 104_1_D0 0.874
#> 5 104_1_D2 104_1_D1 0.109
#> 6 104_1_D2 104_1_D10 0.904R Markdown helpers for scientific writing
When writing scientific papers with R Markdown, we often find
ourselves using the same knitr chunk options and miscellaneous helper
functions. To use our favorite options like eval=TRUE,
echo=FALSE, and others, run set_knitr_opts()
in the first chunk of your R Markdown document:
```{r, include = FALSE}
set_knitr_opts()
```This also sets the inline hook to our custom
inline_hook() function, which automatically formats numbers
in a human-readable way and inserts an Oxford comma into lists when
needed.
Who doesn’t love an Oxford comma?
When writing with R Markdown, you may wish to insert a list or vector
inline and correctly format it with an Oxford comma.
inline_hook() uses paste_oxford_list() to help
you do just that!
animals <- c("cats", "dogs", "fish")Insert the string as inline code with `r `:
`r animals`are the most common pets.
Rendered output:
cats, dogs, and fish are the most common pets.
Human-readable numbers
inline_hook() uses format_numbers() under
the hood to automatically format numbers to a human-readable format,
rather than display in scientific notation.
The numbers
`r c(1e-04, 1e-05, 1e-06)`are very precise, while`r c(1e04, 1e05, 1e06)`are very large.
Rendered output:
The numbers 0.0001, 0.00001, and 0.000001 are very precise. while 10,000, 100,000, and 1,000,000 are very large.
